 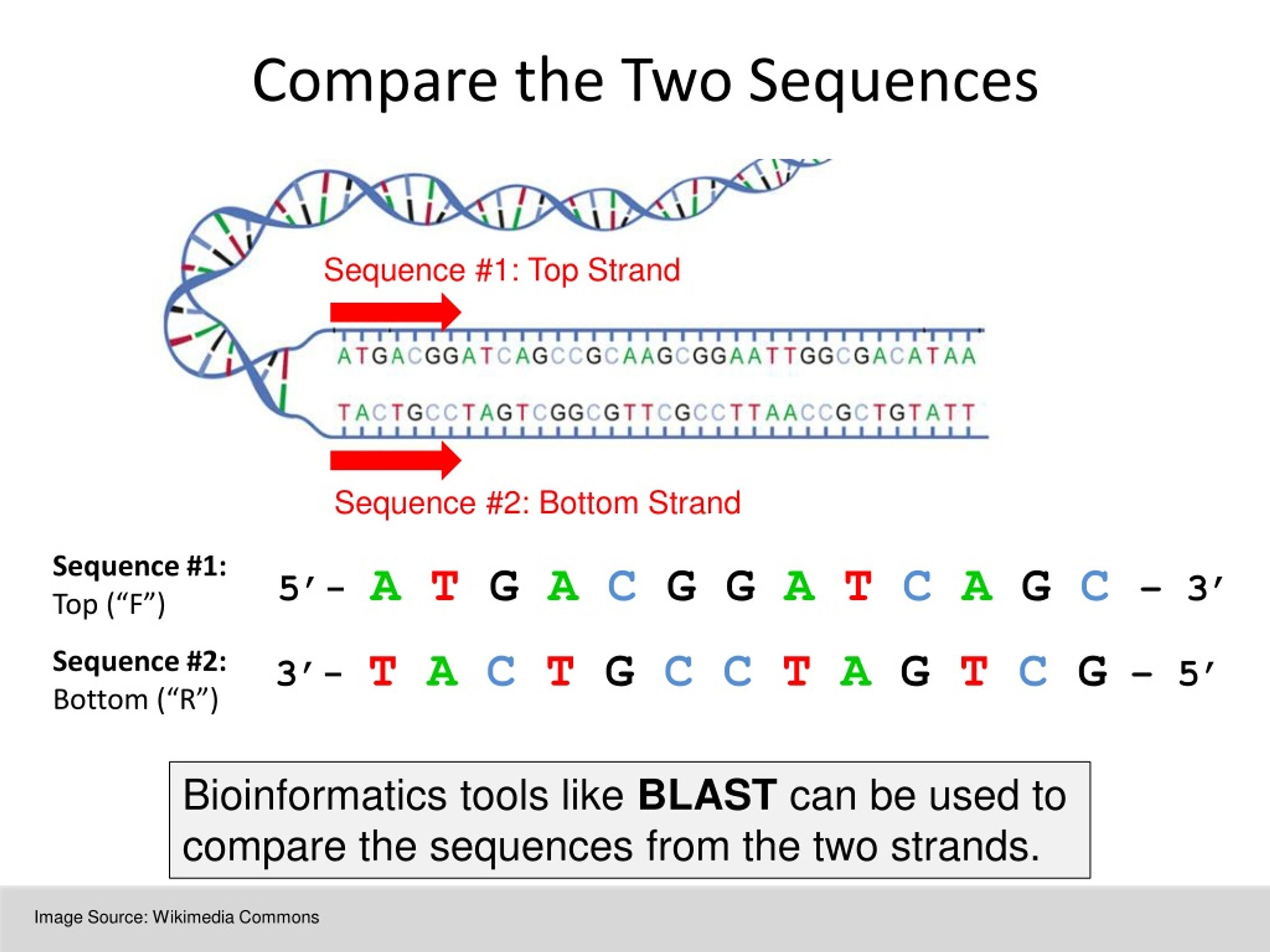
In the above case we are merely lining up the sequences of DNA pairwise, and highlighting the matches between the sequences. An example of performing non-optimal global alignment on our sequences is the following: We’re going to use a scoring method (see below) in conjunction with the Needlman-Wunsch algorithm to figure out the optimal global alignment. We need a metric to use for computing the global alignment between DNA strands. 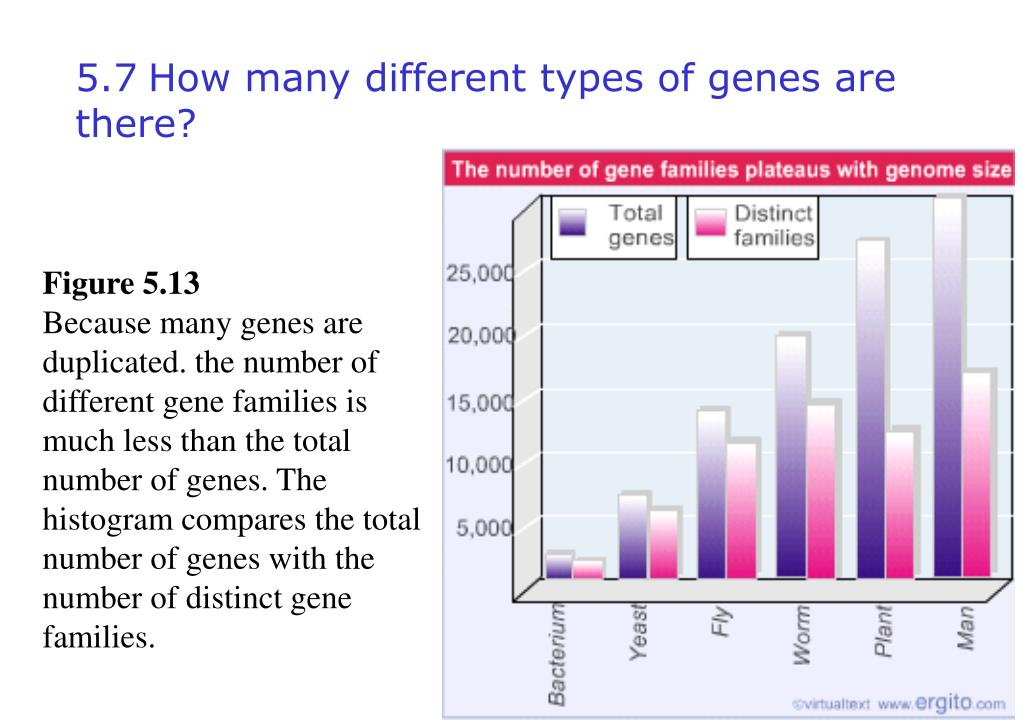
How to measure the similarity between DNA strands Suppose, then, that we have the two sequences of DNA as seen below. Programmatically, DNA can be represented as a string of characters, where each character must be one of A, G, C, or T. These are typically abbreviated by the letters A, G, C, and T, respectively. There are other ways of assessing DNA similarity, and I may cover some of them in a future article, but for now we’ll stick with global alignment.ĭNA is made of four types of nucleotides – adenine, guanine, cytosine, and thymine. By aligning the DNA sequences pairwise, we can create gaps in between nucleotides to create better alignments. “Alignment” means we are trying to align the sequences of DNA such that some score is maximized based upon how many matches / mismatches / gaps we have between the strands. “Global” means we will use the entirety of each sequence of DNA. Backgroundįor our similarity example, we will measure the global alignment between two strands of DNA, and then show how this can be done more generally with dynamic programming. This helps to give insight into the structure and function of a newly identified gene or DNA sequence. 
comparing the DNA of two different species, or two different genes). Measuring the similarity between two different sequences of DNA is very useful because it can help tell us how closely related (or not) those sequences of DNA and their sources are (e.g. Let’s dive into the main topic of this post by implementing an algorithm to measure similarity between two strands of DNA. Click here to read more about dynamic programming. In this way, recursive problems (like the Fibonacci sequence for example) can be programmed much more efficiently because dynamic programming allows you to avoid duplicate (and hence, wasteful) calculations in your code. For anyone less familiar, dynamic programming is a coding paradigm that solves recursive problems by breaking them down into sub-problems using some type of data structure to store the sub-problem results. Dynamic Programming and DNAĭynamic programming has many uses, including identifying the similarity between two different strands of DNA or RNA, protein alignment, and in various other applications in bioinformatics (in addition to many other fields). Here wehave another pair of sequences which are less similar becausethey are three differences. That's quite nice, it'sa hamming distance. Is it really a distance? A distance is a mathematicalconcept and to be a distance, it must satisfy three conditions:the distance between a sequence and itself must be zero, a sequencebetween a sequence and another one must be the same betweenthe last one and the first one and we must have this inequalitywhich is always verified.*Note, if you want to skip the background / alignment calculations and go straight to where the code begins, just click here. So thequestion now is how can we measure this similarity between twosequences for the moment. The first approach to similarityis a very simple one is to apply a distance which is calledhere the Editing System or the Hamming Distance.The idea is very basic. You would take two sequences likethese two sequences here and you look at the differences and youcount the number of differences. Here, for example, you have twodifferences so you will say that the distance, the similaritybetween the two sequences, the distance is two. It's because they evolve togetherwith the species and they evolve in time, there aremodifications in the sequence and that the sequence may still besimilar, similar enough again to retrieve information on onesequence to transfer it to another sequence of interest. So we understand why gene orprotein sequences may be similar. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
